The Polymerase Chain Reaction (PCR) is the workhorse of biotechnology. Its development has revolutionized genetic analysis and engineering science, and many advanced experimental techniques rely on nucleic acid amplification, often in a preprocessing step. In a PCR reaction, DNA is amplified in vitro using an enzymatic reaction. Short DNA sequences complementary to the DNA region to amplify are required to initiate the polymerase reaction, these so-called primers bind to the template DNA, and the synthesis of new DNA begins at these primers.
Multiplexing the Polymerase Chain Reaction, thus amplifying several targets at different genomic locations is of considerable interest, since it can lead to significant time savings and cost reduction. However, to sucessfully multiplex PCR reactions, careful experimental design is necessary.
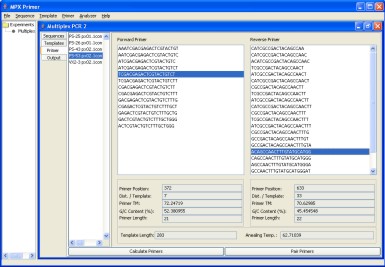
We have developed a Java Computer Program to aid in the primer design process for Multiplex PCR reactions. The software is available for free from this webpage. The software provides a convenient graphical user interface to aid in the experiment design process.

The software is available for free download from the following link. It has been written in the Java Programming Language and should run under any system with a Java Virtual Machine. Java is available from Sun Microsystems and needs to be installed separately.
Download software (zip archive)
Documentation:")?> Documentation and Examples are included in the zip archive available for download above. Contact:")?>Lars Kaderali